A Predicament
Recently we spent some time contributing to dib-lab/eelpond (renamed to elvers), an executable Snakemake workflow for running the eelpond mRNAseq workflow.
In the process of tracking down a confusing bug in the Snakemake workflow, we used Snakemake's ability to print a directed acyclic graph (hereafter referred to as a dag) representing its task graph. Snakemake prints the dot notation to stdout.
(The graph representation ended up identifying the problem, which was two task graphs that were independent, but which were not supposed to be independent.)
When creating the Graphviz dot notation,
Snakemake is kind enough to direct all of its output
messages to stderr, and direct the dot graph output
to stdout, which makes it easy to redirect stdout
to a .dot file and process it with Graphviz.
Github user @bluegenes
(the principal author of elvers) added a --dag file to the run_eelpond script,
which asks Snakemake to print the dag when it calls
the Snakemake API:
./run_eelpond --dag ... > eelpond_dag.dot
This .dot file can then be rendered into a .png file with another command from the command line,
dot eelpond_dag.dot -Tpng -o eelpond_dag.png
Simple, right?
But here's the problem: While this is a simple and easy way to generate the dag, it introduces some extra steps for the user, and it also prevents us from being able to print anything to stdout before or after the dag is generated, since anything printed out by the program to stdout will be redirected to the final dot file along with all the Graphviz dot output.
So how to avoid the extra steps on the command line, while also improving the flexibility in printing to stdout (i.e., only capturing Snakemake's output to a file)?
Can we add two flags like --dagfile and --dagpng
that would, respectively, save the task graph
directly into a .dot file, or render the dot output
from Snakemake directly into a png using dot?
We implemented precisely this functionality in dib-lab/eelpond PR #73. To do this, we utilized a context manager to capture output from Snakemake. In this post we'll cover how this context manager works, and mention a few other possibilities with context managers.
What is a context manager?
If you have done even a little Python programming,
you have probably seen and used context managers
before - they are blocks of code that are
opened using a with keyword in Python. For example,
the classic Pythonic way to write to a file uses
a context manager:
with open('file.txt', 'w') as f:
f.write("\n".join(range(10)))
The context manager defines a runtime context for
all the code in the block - and that can be a different
context than the rest of the program. When a context
is opened (when the with block is encountered),
a context manager object is created and its __enter__()
method is run. This method will modify the runtime
context in whatever way it needs,
and the rest of the code in the block will be run.
When the context is done, where the block ends,
the context manager's __exit__() method is run.
This restores the runtime context to its
normal state for the rest of the program.
It's a general concept with a lot of different
applications. We cover how to use it to capture
output to sys.stdout below.
What is Graphviz dot?
We mentioned that Snakemake can output visualizations of workflows in Graphviz dot format. For the purposes of clarity we explain what that format is here.
Without getting too sidetracked, Graphviz dot defines a notation for drawing graphs, and provides software for laying out the graphs in rendered images.
The user specifies the nodes and labels and edges, as well as formatting and layout details, and dot takes care of laying out the graph.
Here's an example of a simple graph in dot notation:
plot.dot
digraph G {
Boston
"New York"
Houston
"Los Angeles"
Seattle
Boston -> "New York"
Boston -> Houston
Houston -> Boston
Houston -> "Los Angeles"
"New York" -> Seattle
Seattle -> "New York"
"New York" -> "Los Angeles"
}
To render this as a .png image,
dot cities.dot -Tpng -o cities.png
which becomes:
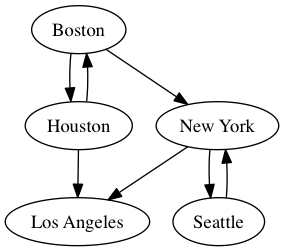
This tool makes visualizing workflows a breeze, as the flow of tasks is much easier to understand and troubleshoot than the convoluted logic of Snakefile rules. Here is an example from elvers:
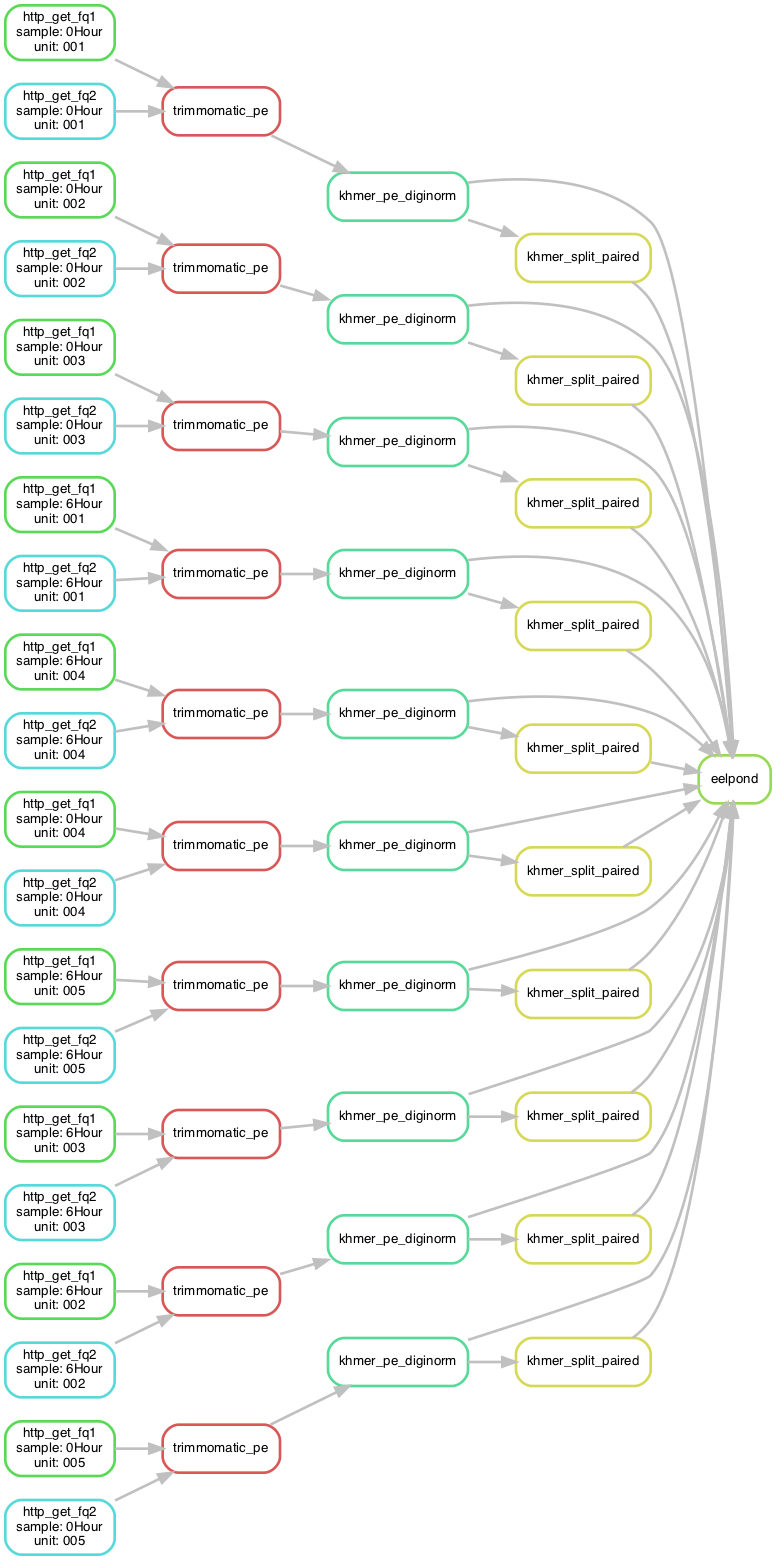
Capturing stdout
In elvers, the run_eelpond command line wrapper that kicks
off the workflow is a Python script that calls the Snakemake
API (we covered this approach in a prior blog post,
Building Snakemake Command Line Wrappers for Workflows).
This Python script has a call to the Snakemake API; here is the relevant snippet:
# ...set up...
if not building_dag:
print('--------')
print('details!')
print('\tsnakefile: {}'.format(snakefile))
print('\tconfig: {}'.format(configfile))
print('\tparams: {}'.format(paramsfile))
print('\ttargets: {}'.format(repr(targs)))
print('\treport: {}'.format(repr(reportfile)))
print('--------')
# Begin snakemake API call
status = snakemake.snakemake(snakefile, configfile=paramsfile, use_conda=True,
targets=['eelpond'], printshellcmds=True,
cores=args.threads, cleanup_conda= args.cleanup_conda,
dryrun=args.dry_run, lock=not args.nolock,
unlock=args.unlock,
verbose=args.verbose, debug_dag=args.debug,
conda_prefix=args.conda_prefix,
create_envs_only=args.create_envs_only,
restart_times=args.restart_times,
printdag=building_dag, keepgoing=args.keep_going,
forcetargets=args.forcetargets,forceall=args.forceall)
# End snakemake API call
# ...clean up...
Most of the code that comes before this API call is processing the flags provided by the user. We want to have the flexibility to print to stdout while processing flags, before we get to the Snakemake API call; and we want those messages to be kept separate from the dag output.
In other words, we only want to capture output to stdout between "Begin snakemake API call" and "End snakemake API call". Everywhere else, stdout can go to stdout like normal.
We can do this by recognizing that any Python program printing to
stdout uses sys.stdout under the hood to send output to stdout -
so if we can somehow tell Python to swap out stdout with a string
buffer that has the same methods (print, printf, etc.), run Snakemake,
then replace stdout again, we can isolate and capture all stdout from
the Snakemake API call.
Replacing stdout
The strategy for our context manager and the entry and exit methods, then, is clear:
-
If the user has specified the
--dagflag, the__entry__()method should replace stdout with a StringIO buffer within our new runtime context; otherwise, leave stdout alone. -
If the user has specified the
--dagflag, the__exit__()method should clean up by restoringsys.stdout; otherwise, do nothing.
Now we are ready to make our context manager object.
But wait! What kind of object are we using? Do we need some kind of special context manager class? Nope! This is one of the features of context managers that makes them magical:
Any object can be a context manager.
All we need to do is add __enter__() and __exit__()
methods to an object, and it can become a context
manager.
Creating a context manager
In our case, we are capturing stdout from Snakemake
so that we can potentially process it, and then dump
it to a file. We don't know how many lines Snakemake
will output, so we will replace sys.stdout with a string
buffer. But once the context closes, we want all those
strings in something more convenient, like a list.
So, we can define a new class that derives from the
list class, and just adds __enter__() and __exit__()
methods, to enable this list to be a context manager:
class CaptureStdout(list):
"""
A utility object that uses a context manager
to capture stdout from Snakemake. Useful when
creating the directed acyclic graph.
"""
def __init__(self,*args,**kwargs):
pass
def __enter__(self,*args,**kwargs):
pass
def __exit__(self,*args,**kwargs):
...
(Note that we include the constructor, since we need the context manager to have a state so that we can restore the original runtime context to the way it was when we're done.)
Constructor
The constructor is where we process any input arguments passed in when the context is created.
Given that we want our context manager to handle the case of a directed acyclic graph by capturing stdout, and do nothing otherwise, we should have a flag in the constructor indicating whether we want to pass stdout through, or whether we want to capture it.
Additionally, we don't need to call the parent
(super) class constructor, i.e., the list constructor,
because we always start with an empty list.
No need to call super().__init__().
Here is the constructor:
class CaptureStdout(list):
"""
A utility object that uses a context manager
to capture stdout from Snakemake. Useful when
creating the directed acyclic graph.
"""
def __init__(self,passthru=False):
# Boolean: should we pass everything through to stdout?
# (this object is only functional if passthru is False)
super().__init__()
self.passthru = passthru
Enter method
When we open the context, we want to swap out
sys.stdout with a string buffer. But we also
want to save the original sys.stdout object
reference, so that we can restore the original
runtime context and let the program continue
printing to stdout after Snakemake is done.
from io import StringIO
import sys
class CaptureStdout(list):
...
def __enter__(self):
"""
Open a new context with this CaptureStdout
object. This happens when we say
"with CaptureStdout() as output:"
"""
# If we are just passing input on to output, pass thru
if self.passthru:
return self
# Otherwise, we want to swap out sys.stdout with
# a StringIO object that will save stdout.
#
# Save the existing stdout object so we can
# restore it when we're done
self._stdout = sys.stdout
# Now swap out stdout
sys.stdout = self._stringio = StringIO()
return self
Exit method
To clean up, we will need to restore sys.stdout
using the pointer we saved in __enter__, then
process the string buffer.
We can also use the del operator to clean up
the space used by the buffer object once we've
transferred its contents.
from io import StringIO
import sys
class CaptureStdout(list):
...
def __exit__(self, *args):
"""
Close the context and clean up.
The *args are needed in case there is an
exception (we don't deal with those here).
"""
# If we are just passing input on to output, pass thru
if self.passthru:
return self
# This entire class extends the list class,
# so we call self.extend() to add a list to
# the end of self (in this case, all the new
# lines from our StringIO object).
self.extend(self._stringio.getvalue().splitlines())
# Clean up (if this is missing, the garbage collector
# will eventually take care of this...)
del self._stringio
# Clean up by setting sys.stdout back to what
# it was before we opened up this context.
sys.stdout = self._stdout
In action
To see the context manager in action, let's go back to the snippet of code where we call the Snakemake API:
# Set up a context manager to capture stdout if we're building
# a directed acyclic graph (which prints the graph in dot format
# to stdout instead of to a file).
# If we are not bulding a dag, pass all output straight to stdout
# without capturing any of it.
passthru = not building_dag
with CaptureStdout(passthru=passthru) as output:
# run!!
# params file becomes snakemake configfile
status = snakemake.snakemake(snakefile, configfile=paramsfile, use_conda=True,
targets=['eelpond'], printshellcmds=True,
cores=args.threads, cleanup_conda= args.cleanup_conda,
dryrun=args.dry_run, lock=not args.nolock,
unlock=args.unlock,
verbose=args.verbose, debug_dag=args.debug,
conda_prefix=args.conda_prefix,
create_envs_only=args.create_envs_only,
restart_times=args.restart_times,
printdag=building_dag, keepgoing=args.keep_going,
forcetargets=args.forcetargets,forceall=args.forceall)
Once we have closed the runtime context, our variable
output is a list with all the output from Snakemake
(assuming we're creating a dag; if not, everything is
passed through to stdout like normal).
The last bit here is to handle the three different
dag flags: --dag, --dagfile, and --dagpng.
-
--dagprints the dot graph straight to stdout, like Snakemake's default dag behavior; -
--dagfile=<dotfile>dumps the dot graph to a dot file -
--dagpng=<pngfile>uses dot (installed in the elvers conda environment) to render the dot output from Snakemake directly into a png image
We handle these three cases like so:
if building_dag:
# These three --build args are mutually exclusive,
# and are checked in order of precedence (hi to low):
# --dag to stdout
# --dagfile to .dot
# --dagpng to .png
if args.dag:
# straight to stdout
print("\n".join(output))
elif args.dagfile:
with open(args.dagfile,'w') as f:
f.write("\n".join(output))
print(f"\tPrinted workflow dag to dot file {args.dagfile}\n\n ")
elif args.dagpng:
# dump dot output to temporary dot file
with open('.temp.dot','w') as f:
f.write("\n".join(output))
subprocess.call(['dot','-Tpng','.temp.dot','-o',args.dagpng])
subprocess.call(['rm','-f','.temp.dot'])
print(f"\tPrinted workflow dag to png file {args.dagpng}\n\n ")
Note that before the Snakemake API call, we also check
whether dot exists:
# if user specified --dagpng,
# graphviz dot must be present
if args.dagpng:
if shutil.which('dot') is None:
sys.stderr.write(f"\n\tError: Cannot find 'dot' utility, but --dotpng flag was specified. Fix this by installing graphviz dot.\n\n")
sys.exit(-1)
The final code is implemented in the cmr_better_dag_handling branch of eelpond/elvers
and pull request #73 in eelpond/elvers.
Using the new dag flags
$ git clone https://github.com/dib-lab/eelpond.git
$ cd eelpond
$ conda env create --file environment.yml -n eelpond
$ conda activate eelpond
Now we're ready to run the workflow.
We can use the -w flag to list all workflows,
then use the -n flag to do a dry run.
We'll use the kmer_trim workflow target:
$ ./run_eelpond examples/nema.yaml kmer_trim -n
...lots of output...
Job counts:
count jobs
1 eelpond
10 http_get_fq1
10 http_get_fq2
10 khmer_pe_diginorm
10 khmer_split_paired
10 trimmomatic_pe
51
This was a dry-run (flag -n). The order of jobs does not
reflect the order of execution.
Now we can create a dag for this workflow target:
$ ./run_eelpond examples/nema.yaml kmer_trim --dagfile=dag_kmertrimming.dot
Added default parameters from rule-specific params files.
Writing full params to examples/.ep_nema.yaml
Building DAG of jobs...
Printed workflow dag to dot file dag_kmertrimming.dot
Finally, we can use the --dagpng flag for instant
gratification:
$ ./run_eelpond examples/nema.yaml kmer_trim --dagpng=dag_kmertrimming.png
Added default parameters from rule-specific params files.
Writing full params to examples/.ep_nema.yaml
Building DAG of jobs...
Printed workflow dag to png file dag_kmertrimming.png
Note that you can add a line rankdir=LR; to your dot
file to change the orientation of the graph (left-to-right
order makes highly-parallel workflows vertically stretched,
so they are eaiser to view).
digraph mydigraph {
rankdir=LR;
...
and here is the result:

Other context manager applications
Actions requiring temporary contexts, which are a bit like self-contained workspaces, are good candidates for context managers. Following are a few examples and references.
SSH connections: the context manager's __enter__ function creates/loads
connection details, creates a connection object, and opens the connection.
The __exit__ function cleans up by closing the connection. This way,
you can say something like
with SSHConnectionManager(device) as conn:
try:
conn.send_command("echo hello world")
except Exception as e:
print("Enountered an error running remote command", e)
Blog post: Using Python Context Managers for SSH connections
(Note: this blog post uses a context manager that is a generator decorated with a context manager utility function; this is a different approach than our class-based approach but is still valid.
iPython notebook and matplotlib figure management: Camille Scott has a blog post covering a way of managing large Jupyter notebooks with lots of figures using context managers. In this case, the context that is being set up and torn down is a matplotlib plot context, and it is creating each plot, saving it to a file, then closing the plot. This makes the notebook a lot faster than trying to render every single plot, and makes it a lot cleaner than littering the code with manual figure and axis management.
Here is the example usage Camille gives:
with FigManager('genes_per_sample', figsize=tall_size) as (fig, ax):
genes_support_df.sum().plot(kind='barh', fontsize=14,
color=labels_df.color,
figure=fig, ax=ax)
ax.set_title('Represented Genes per Sample')
FileLink('genes_per_sample.svg')
(The FileLink function opens/processes the resulting image.)
Nice!
Blog post: Context Managers and IPython Notebook
Database Connection: this example comes from Django's test suite. In it, a context manager is defined that creates new MySQL database connections when the context is opened, and closes them when the context is done.
Here is the context manager, which again uses the decorator + generator approach rather than the class approach:
django/tests/backends/mysql/tests.py:
@contextmanager
def get_connection():
new_connection = connection.copy()
yield new_connection
new_connection.close()
The first two lines of this function are equivalent to an __enter__ method,
while the last line is equivalent to an __exit__ method.
This context manager is then used like so:
with get_connection() as new_connection:
new_connection.settings_dict['OPTIONS']['isolation_level'] = self.other_isolation_level
self.assertEqual(
self.get_isolation_level(new_connection),
self.isolation_values[self.other_isolation_level]
)
The context manager is a convenient way of creating a copy of an existing MySQL connection, then closing it when the requesting method is finished using it.
References
-
- this mRNAseq data processing protocol served as the original inspiration for elvers
-
Building Snakemake Command Line Wrappers for Workflows (charlesreid1 blog)